Alexis has also doe some work on phylogeny visualisation, including using treemaps. See (doi:10.1007/11573067_29, PDF here).
Rants, raves (and occasionally considered opinions) on phyloinformatics, taxonomy, and biodiversity informatics. For more ranty and less considered opinions, see my Twitter feed.
ISSN 2051-8188. Written content on this site is licensed under a Creative Commons Attribution 4.0 International license.
Thursday, August 30, 2007
RAxML BlackBox
Alexis Stamatakis and Jacques Rougemont have released RAxML BlackBox, a prototype Web-Server for RAxML which is attached to a 200 CPU-cluster located at the Vital-IT unit of the Swiss Institute of Bioinformatics. You can upload your data and the cluster will mul it over for up to 24 hours, so typically you can analyse alignments up to 1,000 to 1,500 sequences at present. The software itself can handle bigger data sets (see the RAxML page at CIPRES).
Alexis has also doe some work on phylogeny visualisation, including using treemaps. See (doi:10.1007/11573067_29, PDF here).

Alexis has also doe some work on phylogeny visualisation, including using treemaps. See (doi:10.1007/11573067_29, PDF here).
Monday, August 20, 2007
Internet Explorer -- argh!!!!!
I would prefer to avoid Microsoft-bashing, but today I've spent time trying to get my tree viewer to work under Internet Explorer 6 and 7, and it's hell. Here are the problems I've had to deal with:
Empty DIV bug
On IE 6 the top of the scrollbar overlapped the transparent area when the page first loads. Eventually discovered that this is a bug in IE. It gives empty DIVs a height approximately equal to the font-height for the DIV, even if the DIV has height:0px; (see here for a discussion). I set the CSS for this DIV to overflow:hidden;, and the DIV now behaves.
Opacity
The viewer makes use of opacity, that is, having DIVs that are coloured, but which you can see through. This enables me to add layers over the top of an image. IE doesn't support the standard way of doing this, so styles such as opacity:0.5; must also be written as filter:alpha(opacity=50); (thanks to David Shorthouse for pointing this out to me).
Background transparency
The DIV overlaying the big tree has background-color:transparent;, which means it refuses to accept any mouse clicks on the big tree. Changing the color to anything else meant the DIV received the clicks, so I ended up using a fairly ugly hack to include Internet Explorer-specific CSS for this DIV (idea borrowed from How to Use Different CSS Style Sheets For Different Browsers (and How to Hide CSS Code from Older Browsers)).
z-index bug
The final show stopper was the auto-complete drop down list of taxon names. On IE it disappeared behind the big tree. This is the infamous z-index bug. The drop down menu is a DIV created on the fly, and although it's z-index value (99) means it should be placed on top of the tree (so the user can see the list of taxa), it isn't. After some Googling I settled on the hack of setting the z-index for the DIV containing the big tree to -1 in IE only, and this seems to work.
IE is sometimes good
Sometimes it has to be said, IE has it's good points. The tree viewer failed in part because I'd failed to define a Javascript variable. Somehow FireFox and Safari were OK with this, but IE 6 broke. I defined the variable and it worked. I've also learnt to avoid some variable names, such as scroll. I find FireFox to a better browser for developing stuff, especially if the Firebug extension is installed. However, the Internet Explorer Developer Toolbar is useful if you need to figure out what IE is doing.
Conclusion
It's staggering how much time one can waste trying to cater to the weird and wonderful ways of Internet Explorer. However, the tree viewer should now work for those of you running Internet Explorer.
Saturday, August 18, 2007
Visualising very big trees, Part V
Inspired partly by the image viewers mentioned earlier, and tools like Google Finance's plot of stock prices, I've built yet another demo of one way to view large trees.

You can view the demo here. On the left is a thumbnail of the tree, on the right is the tree displayed "full scale", that is, you can read the labels of every leaf. In the middle appears a subset of any internal node labels. Top right is a text box in which you can search for a taxon in the tree.
You can navigate by dragging the scroll bar on the left, dragging the big tree, or using the mouse wheel (and you can jump to a taxon by name). It has been "tested" in Safari and Firefox on a Mac, I doubt it works on Internet Explorer. Getting that to happen is a whole other project.
The viewer is written entirely in HTML and Javascript, the underlying tree images (and some of the HTML and Javascript) are generated using a C++program that reads and draws trees, and I use ImageMagick to generate actaual images.

You can view the demo here. On the left is a thumbnail of the tree, on the right is the tree displayed "full scale", that is, you can read the labels of every leaf. In the middle appears a subset of any internal node labels. Top right is a text box in which you can search for a taxon in the tree.
You can navigate by dragging the scroll bar on the left, dragging the big tree, or using the mouse wheel (and you can jump to a taxon by name). It has been "tested" in Safari and Firefox on a Mac, I doubt it works on Internet Explorer. Getting that to happen is a whole other project.
The viewer is written entirely in HTML and Javascript, the underlying tree images (and some of the HTML and Javascript) are generated using a C++program that reads and draws trees, and I use ImageMagick to generate actaual images.
Friday, August 17, 2007
Bird supertree project - "Open Source" phylogenetics
Today is the day Katie Davis and I are launching the Bird Supertree Project. Partly an effort to distribute the task of building the tree, partly an experiment in "open source phylogenetics", we're curious (if not anxious) to see how this works out. We encourage anybody who is interested in constructing big trees to visit the site, grab the data and have a play. You can upload your results (and see who is the best tree builder), and view the trees using one of the methods for viewing big trees that I mentioned earlier. I'm frantically trying to improve this viewer, so the more trees we get the greater the incentive for me to improve it.
Wednesday, August 15, 2007
Expand-Ahead: A Space-Filling Strategy for Browsing Trees
For the "to do" list, expand-ahead browsing looks like a useful approach to build upon PygmyBrowse (see my live demo). The approach is described in "Expand-Ahead: A Space-Filling Strategy for Browsing Trees" by McGuffin et al. (doi:10.1109/INFOVIS.2004.21, PDF also here).
There is a video on Ravin Balakrishnan's site, which is an AVI file that I haven't bee able to coerce my Mac into playing, hence I've posted it on YouTube.
There is a video on Ravin Balakrishnan's site, which is an AVI file that I haven't bee able to coerce my Mac into playing, hence I've posted it on YouTube.
Saturday, August 11, 2007
Visualising very big trees, Part IV

Continuing the theme of viewing big trees, another approach to viewing large objects is tiling, which most people will have encountered if they've used Google Maps.The idea is to slice a large image into many smaller pieces ("tiles") at different reoslutions, and display only those tiles needed to show the view the user is interested in. I'd thought about doing this for trees but abandoned it. However, I think it is worth revisiting, based on discussion on the Nature Network Bioinformatics Forum, and looking at the Giant-Ass Image Viewer (version 2 is here), and Marc Paschal's blog.
As an example of what could be done, below is a phylogeny from Frost et al.'s "The amphibian tree of life" hdl:2246/5781, rendered using Zoomify's Zoomify Express. I just took a GIF I'd made of the entire tree, dropped it on the Zoomify Express icon, hacked some HTML, and got this:
Now, I don't think Zoomify itself is the answer, because what I'd like is to constrain the navigation to be in one dimension, to have a clearer sense of where I am in the tree, and to have a search function to locate nodes of interest. However, this approach seems worth having a look at. Looks like I'll need to learn a lot more Javascript...
Saturday, August 04, 2007
Visualising very big trees, Part III
I've refined my first efforts to now highlight where you are in the tree. The trees on display here now show the new look.
Basically I've abandoned image maps as they don't allow me to highlight the part of the tree being selected. After some fussing I switched to using HTML DIVs, which sit on top of the image. This took a little while to get working, CSS and DIV placement drives me nuts. The trick is to give each DIV the style position:absolute;, and (and this is important) make sure that the DIV is written as <div ...></div>, not <div .../>.
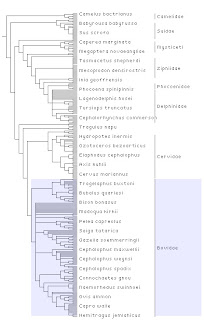
The trees now show a pale blue highlight when you mouse over an area you can click on, and if the corresponding subtree has an internal node label, that label is also highlighted. In the same way, if you mouse over the region on the right that corresponds to a labelled internal node, botht he label and the subtree are highlighted. I think this helps make it clear what parts you are selecting, and gives you the option of selecting using a name, rather than clicking on part of the tree.
Basically I've abandoned image maps as they don't allow me to highlight the part of the tree being selected. After some fussing I switched to using HTML DIVs, which sit on top of the image. This took a little while to get working, CSS and DIV placement drives me nuts. The trick is to give each DIV the style position:absolute;, and (and this is important) make sure that the DIV is written as <div ...></div>, not <div .../>.

The trees now show a pale blue highlight when you mouse over an area you can click on, and if the corresponding subtree has an internal node label, that label is also highlighted. In the same way, if you mouse over the region on the right that corresponds to a labelled internal node, botht he label and the subtree are highlighted. I think this helps make it clear what parts you are selecting, and gives you the option of selecting using a name, rather than clicking on part of the tree.
Friday, August 03, 2007
Visualising very big trees, Part II
OK, time to put my money where my mouth is. Here's a first stab at displaying big trees in a browser. Not terribly sophisticated, but reasonably fast. Take a look at Big Trees.
Approach
Given a tree I simply draw it in a predetermined area (in these examples 400 x 600 pixels). If there are more leaves than can be drawn without overlapping I simply cull the leaf labels. If there are internal node labels I draw vertical lines corresponding to the span of the corresponding subtree, which is simply the range between the left-most and right-most decendants of that node. If internal node labels are nested (e.g., "Mammalia" and "Primates") I draw the most recent internal node label, the rationale being that I want only a single set of vertical bars. This gives the visual effect of partitioning up the leaves into non-overlapping sets. This gives us a diagram like this:
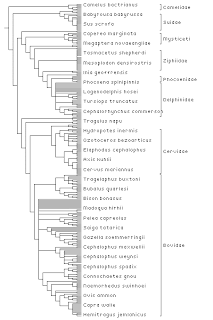
OK, but what about all the nodes we can't see? What I do here is make the tree "clickable" in the following way. If there are internal node labels I make the corresponding tree clickable. I also traverse the tree looking for well defined clusters -- basically subtrees that are isolated by a long branch from their nearest neighbours -- and make these clickable. This approach is partly a hang over form earlier experiments on automatically folding a tree (partly inspired by doi:10.1111/1467-8659.00235). The key point is I'm trying to avoid testing for mouse clicks on nodes and edges, as many of these will be ocluded by other nodes and edges, and it will also be expensive to do hit testing on nodes and edges in a big tree.
If you click on one the script extracts the subtree and reloads the display showing just that part of the tree, using exactly the same approach as above. Behind the scenes the code is doing a least common ancestor (LCA) query, hence it defines subtrees rather like the Phylocode does (oh the irony).
Pros
Cons
Approach
Given a tree I simply draw it in a predetermined area (in these examples 400 x 600 pixels). If there are more leaves than can be drawn without overlapping I simply cull the leaf labels. If there are internal node labels I draw vertical lines corresponding to the span of the corresponding subtree, which is simply the range between the left-most and right-most decendants of that node. If internal node labels are nested (e.g., "Mammalia" and "Primates") I draw the most recent internal node label, the rationale being that I want only a single set of vertical bars. This gives the visual effect of partitioning up the leaves into non-overlapping sets. This gives us a diagram like this:

OK, but what about all the nodes we can't see? What I do here is make the tree "clickable" in the following way. If there are internal node labels I make the corresponding tree clickable. I also traverse the tree looking for well defined clusters -- basically subtrees that are isolated by a long branch from their nearest neighbours -- and make these clickable. This approach is partly a hang over form earlier experiments on automatically folding a tree (partly inspired by doi:10.1111/1467-8659.00235). The key point is I'm trying to avoid testing for mouse clicks on nodes and edges, as many of these will be ocluded by other nodes and edges, and it will also be expensive to do hit testing on nodes and edges in a big tree.
If you click on one the script extracts the subtree and reloads the display showing just that part of the tree, using exactly the same approach as above. Behind the scenes the code is doing a least common ancestor (LCA) query, hence it defines subtrees rather like the Phylocode does (oh the irony).
Pros
- Reasonably fast (everything you see is done live "on the fly").
- Works in any modern browser, no dependence on plugins or technology that has limited support.
- Image is clear, text is small but legible.
- Entirely automated layout
Cons
- Reloading a new page is costly in terms of time, and potentially disorienting (you loose sense of the larger tree).
- It is not obvious where to click on the tree (needs to be highlighted).
- Text is not clickable. This is would be really useful for internal node labels.
Thursday, August 02, 2007
Viewing very large trees

One of the striking pictures in Tamara Munzner et al.'s paper "TreeJuxtaposer: Scalable Tree Comparison using Focus+Context with Guaranteed Visibility" (doi:10.1145/882262.882291, also available here) is that of a biologist struggling to visualise a large phylogeny. The figure caption states that:
Biologists faced with inadequate tools for comparing large trees have fallen back on paper, tape, and highlighter pens.
I've been struggling with this problem in the context of display trees on a web page (see an earlier post). Viewing large trees has received a lot of attention, and there are some fun tools such as Tamara Munzner's TreeJuxtaposer and Mike Sanderson's Paloverde (doi:10.1093/bioinformatics/btl044 ), which was used to create the cover for the October 2006 issue of Systematic Biology. And let's not forget Google Earth.

The problem with standalone tools like these is that they are just that - standalone. They are meant to support interactive visualisation in an application, not viewing a tree on a web page. This is a particular problem facing TreeBASE. A user wanting to view, say, the marsupial supertree published by Cardillo et al. (doi:10.1017/S0952836904005539, TreeBASE study S1035) is greeted by the message:
This tree is too large to be seen using the usual GUI. We recommend that you view the tree using the java applet ATV or the program TreeView (see below). Alternatively, you can download the data matrix and view the tree(s) in MacClade, PAUP, or any other nexus-compatible software.and the tree is displayed as Newick text string:
(((((((((((Abacetus, ((((Agonum, Glyptolenus), Europhilus, Tanystoma, Platynus), ((Morion, Moriosomus), Stenocrepis)), (((Licinus, Zargus, Badister), (Panagaeus, Tefflus)), Melanchiton)), ((Amara, Zabrus), (Harpalus, Dicheirotrichus, Parophonus, Trichocellus, Ophonus, Trichotichnus, Diachromus, Pseudoophonus, Stenolophus, Notobia, Bradycellus, Nesacinopus, Anisodactylus, Acupalpus, Acinopus, Xestonotus)), ((Anthia, Thermophilum), ((Corsyra, Discoptera), Graphipterus)), (((Apenes, (Chlaenius, Callistus)), Oodes), ((Calophaena, ((Ctenodactyla, Leptotrachelus), Galerita)), (Pseudaptinus, Zuphius))), (((Calleida, Hyboptera), Lebia), Cymindis, Demetrias, Dromius, Lionychus, Microlestes, Syntomus), ((Calybe, Lachnophorus), Odacantha), (Catapiesis, (Desera, Drypta)), Cnemalobus, (Coelostomus, (Eripus, Pelecium)), ...
 Not the most compelling visualisation. What I hope to do in this and following posts is describe my own efforts to come to grips with this problem.
Not the most compelling visualisation. What I hope to do in this and following posts is describe my own efforts to come to grips with this problem.Requirements
To put the problem into perspective, what I'm looking for is a simple way to draw large trees for display in a web browser. This places severe limits on the kind of interactivity that is possible (unless we go down the root of Java applets, which I will avoid like the plague). This rules out, for example, trying to emulate TreeJuxtaposer's functionality. Initially I started looking at SVG, which renders graphics nicely, supports interaction, and being essentially an XML file, is easy to manipulate (for an example see my earlier post on SVG maps). However, SVG is not well supported in all browsers (FireFox does pretty well, most other browsers are variable). All browsers, however, support bitmap graphics (GIF, PNG, JPEG, etc.). When drawing complex things like trees bitmaps have some advantages, especially with regards to labelling. Small bitmap fonts tend to be more legible than anti-aliased fonts at the same size (see article at MiniFonts for background.
Animation
Comments so far on this post have focussed on animation (e.g., using Flash). Here is a video of TreeJuxtaposer taken from Tamara Munzner's web site.
For me the most interesting features of TreeJuxtaposer are that the entire tree is always visible (thus retaining context, unlike pan and zoom), and the user can select bits to view by drawing a rectangle on the screen. The processing to compute the transformations needed for large trees is fairly heavy duty, although newer algorithms have reduced this somewhat (see here).
Subscribe to:
Posts (Atom)